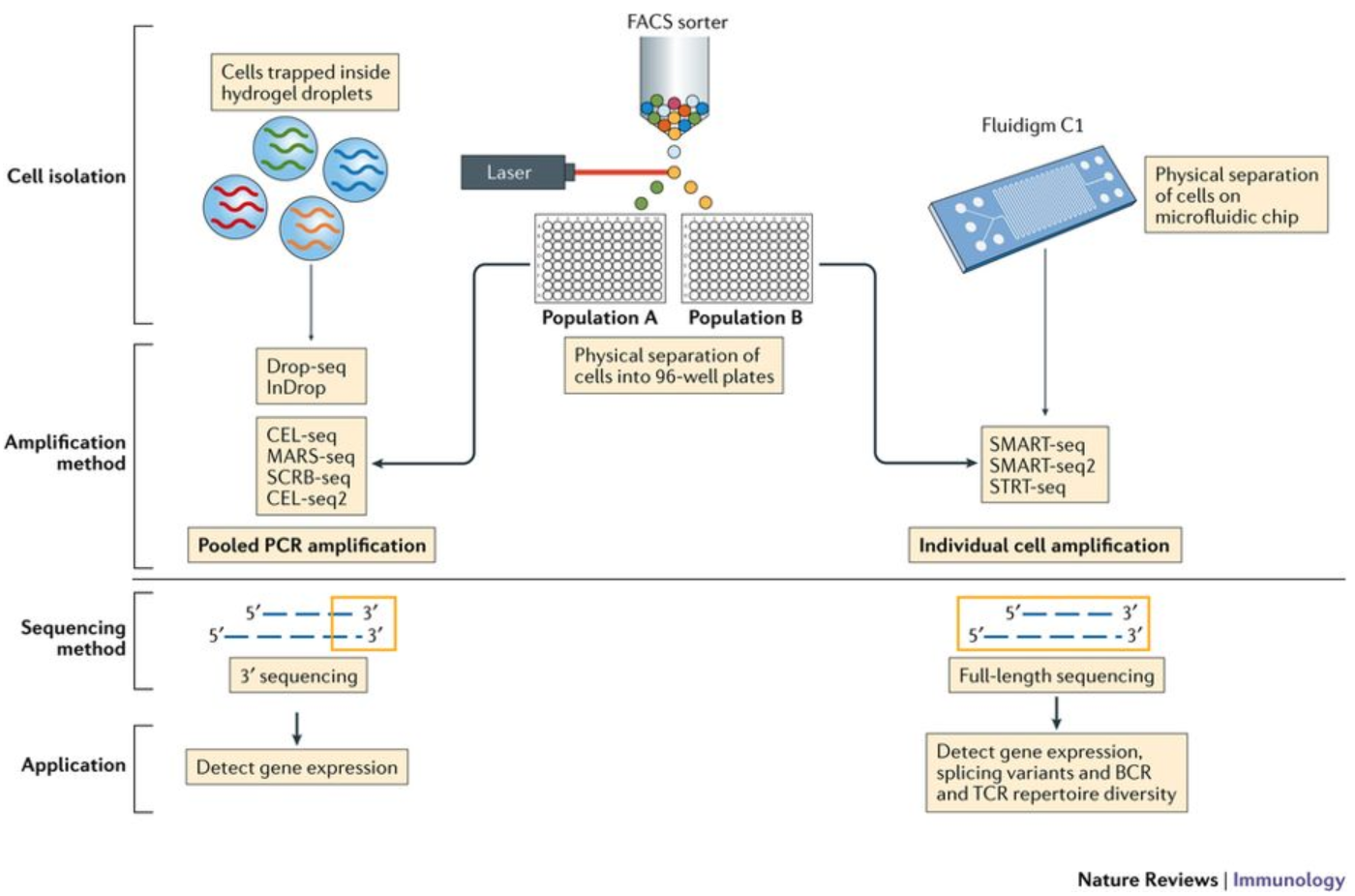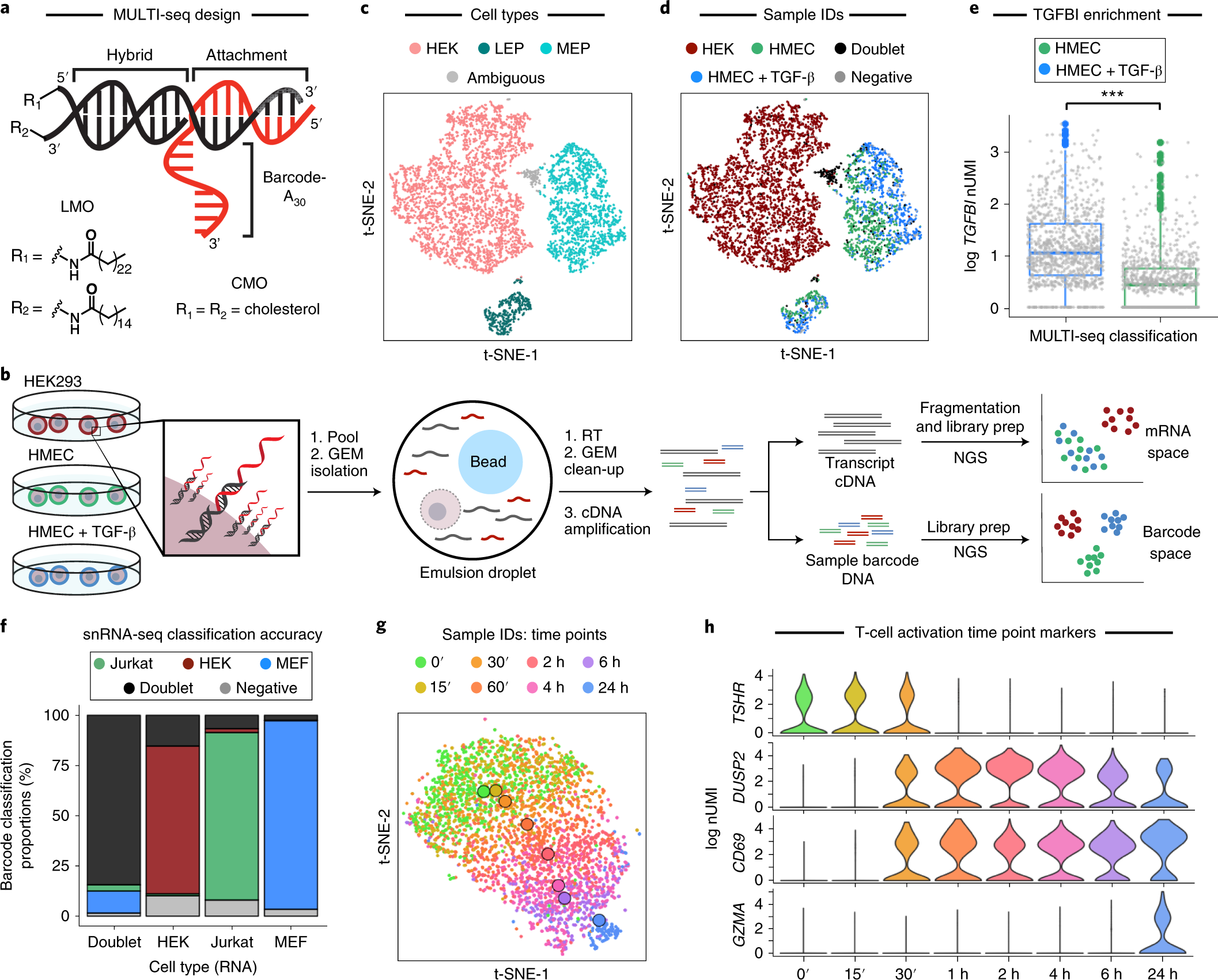
MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices | Nature Methods

Second-Strand Synthesis-Based Massively Parallel scRNA-Seq Reveals Cellular States and Molecular Features of Human Inflammatory Skin Pathologies - ScienceDirect

Principle and workflow of MAPS-seq. a Cells were dispensed into 96-well... | Download Scientific Diagram

Ragon Institute on Twitter: "@Shaleklab and J. Christopher Love, @MITChemE @kochinstitute and Ragon Associate Member, recently published their new single-cell RNA sequencing technique, Seq-Well S3, in @ImmunityCP @CellPressNews @MIT https://t.co ...
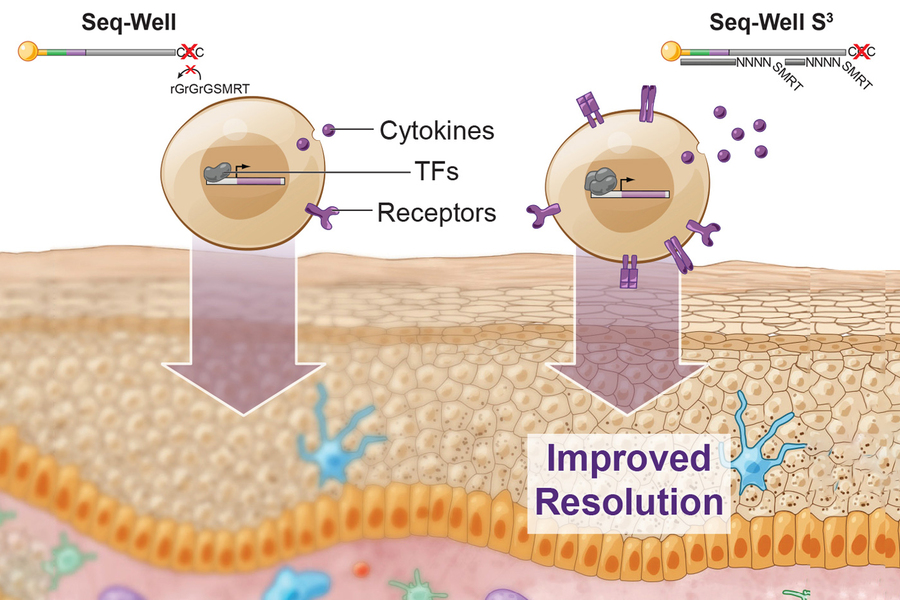
Technique recovers lost single-cell RNA-sequencing information | MIT News | Massachusetts Institute of Technology

Second-Strand Synthesis-Based Massively Parallel scRNA-Seq Reveals Cellular States and Molecular Features of Human Inflammatory Skin Pathologies - ScienceDirect
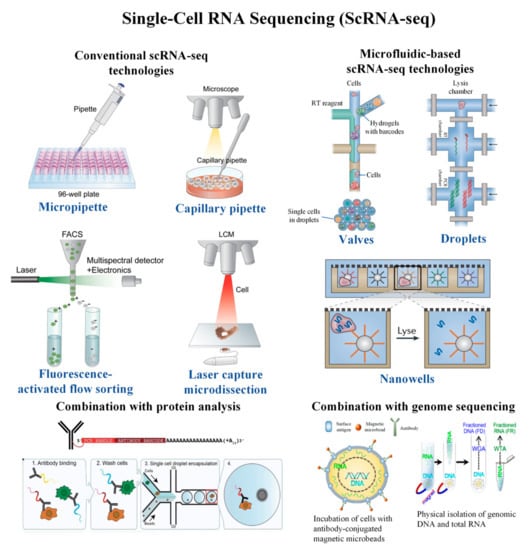
Cells | Free Full-Text | Single-Cell RNA Sequencing and Its Combination with Protein and DNA Analyses
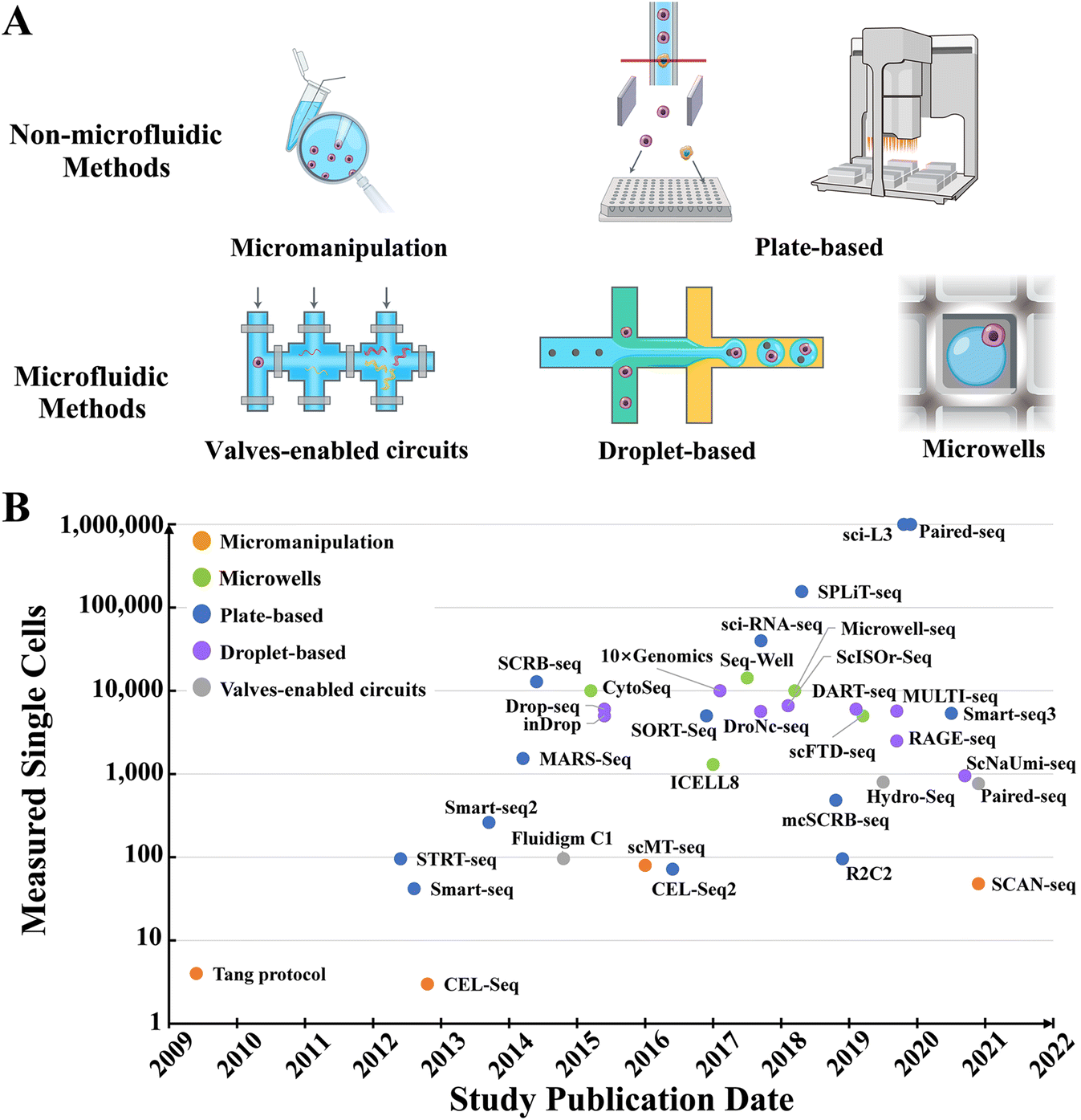
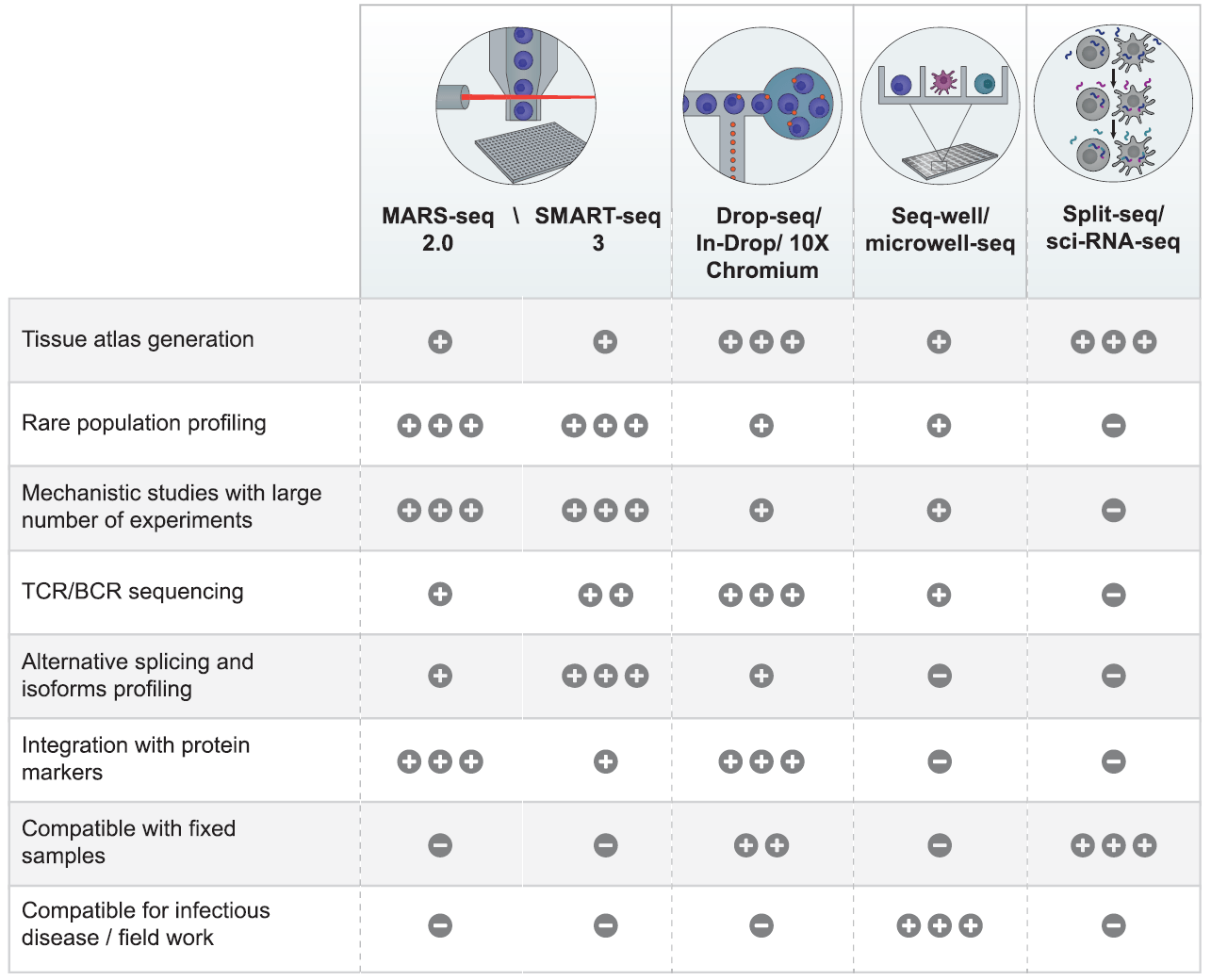
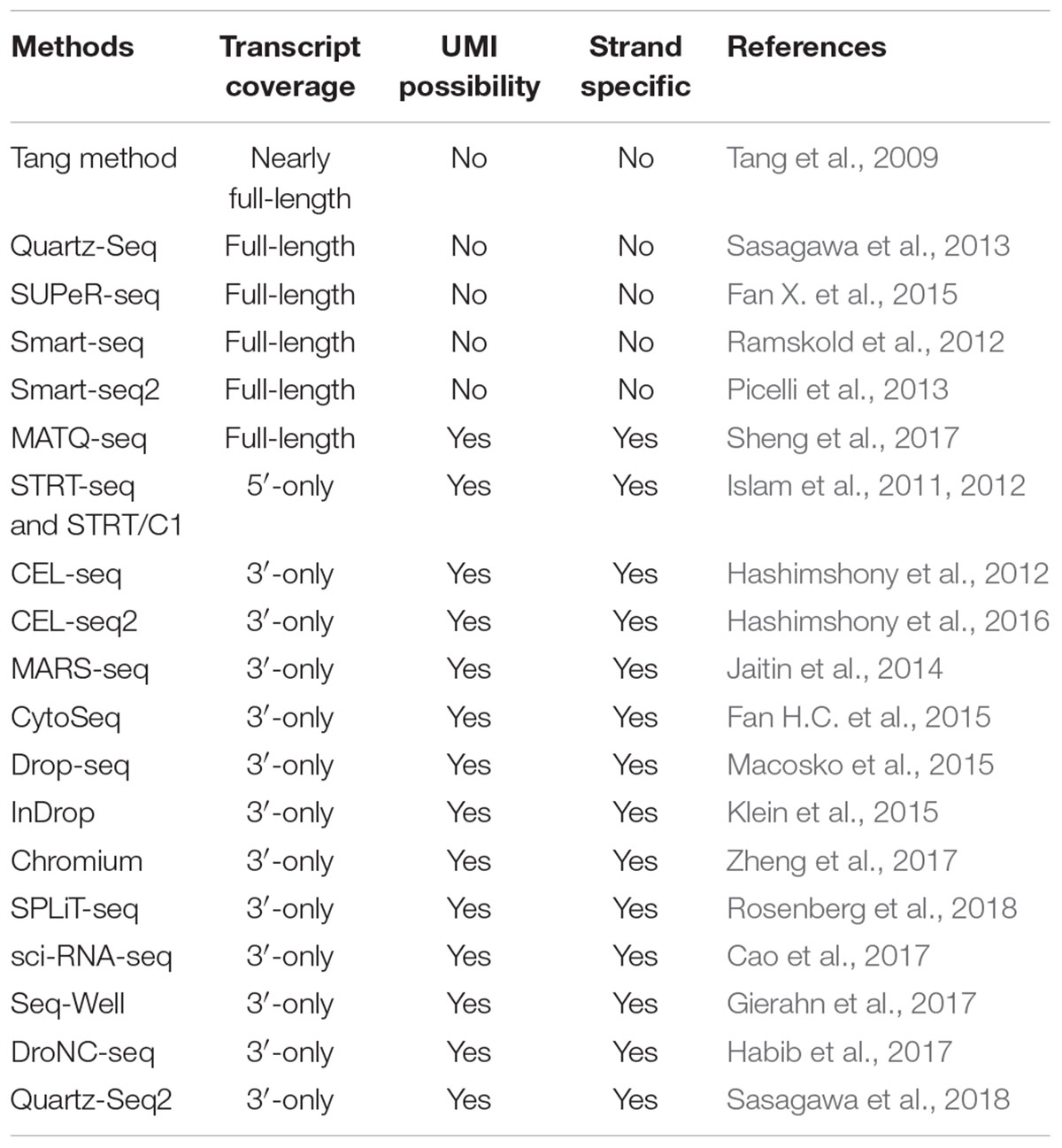
Recent advances in high-throughput single-cell transcriptomics and spatial transcriptomics - Lab on a Chip (RSC Publishing) DOI:10.1039/D2LC00633B

Well-TEMP-seq – a microwell-based strategy for massively parallel profiling of single-cell temporal RNA dynamics | RNA-Seq Blog

Diverse Myeloid Cell States Uncovered by using Seq-Well S 3 (A) (Left)... | Download Scientific Diagram